Python environments or how to survive to your journey in the geodata space
Python is currently the election language for the data science (including the geospatial software ecosystem, both FOSS and proprietary). In respect to other languages it is has a long history of breaking changes with a development cycle of not more than 5 years per version. From time to time you can find changes in tooling and packages, or even big turns in the major versions (e.g. Python 2 >> Python 3 required a 10 years roadmap until the end of 2.* series, with deprecation of a lot of packages and changes in syntax).
Main aspects to consider in your workflow:
Multiple platforms (operating systems) involve different constraints.
Many packages (the most performant) are not native i.e. are not written (completely) in Python. That includes the Big Ones (numpy, pandas, xarray, etc.).
The so-called extension packages are based on external libraries written in C/C++.
Multiple toolchains have been introduced to simplify the workflow on some platforms, but the most advanced use could be tricky.
Documentation is not always complete or articulated enough. In some cases sources or some users forum are the last resort.
[3]:
from IPython.display import Image
Image("../images/pythonrels.png" , width = 800, height = 800)
[3]:
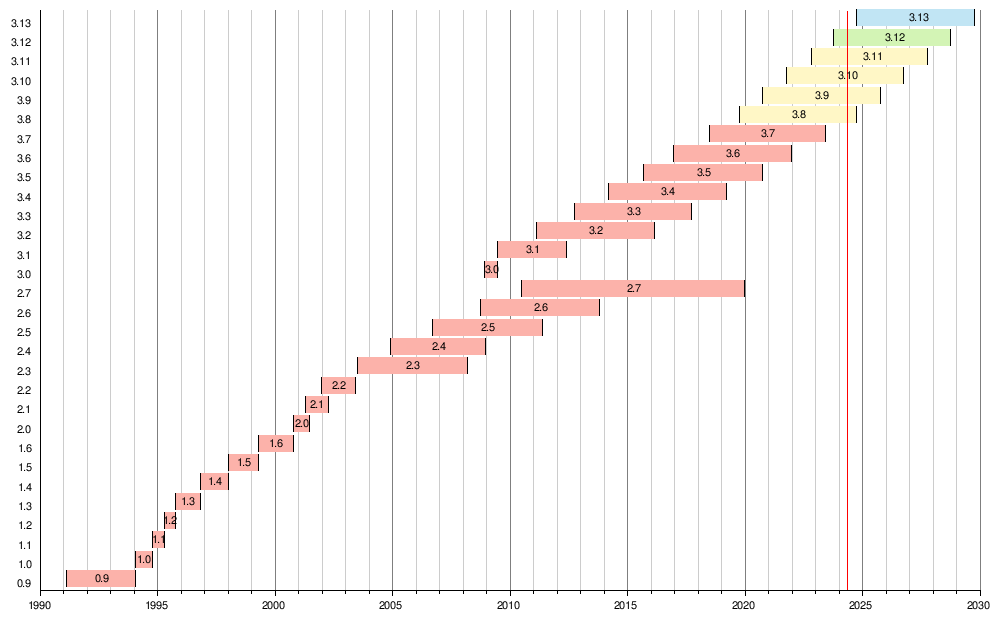
A brief list of distilled suggestions and considerations
Linux and other platforms have a single system Python with pre-built packages available in the distribution archive. They also have a per-release tool chain, including a C/C++ compiler and standard libraries set.
DON’T try to change system versions, you can get a not functional system which could require a total reinstall.
With the right tools any user can install and use multiple Python versions without breaking the system. See PyEnv and Anaconda.
With the right tools any user can install multiple packages versions to use alternatively. See venv environments and Anaconda.
Unfortunately you CAN’T use multiple versions of the same package at the same time. I’m very sorry, guys :-)
All the previous stuff in general do not require admin privileges, so you could use them within scientific clusters too.